Case Study: How MDx Helped Fulgent Genetics Achieve CE Marking for One of the World’s Largest Germline NGS Panels under IVDR
IVDR CE marking NGS at a glance
- Outcome: CE mark granted by TÜV SÜD for an end-to-end Class C germline NGS solution (wet lab + bioinformatics).
- Scope: Furthermore, panel covering 4,600+ clinically relevant genes with a validated PLM (Pipeline Manager) software component; later expanded to >7,000 genes using a new probe set.
- What we did: Specifically, we built an ISO 13485 QMS from the ground up, prepared full Annex II + III technical documentation, validated bioinformatics under IEC 62304/82304, split documentation into two Basic UDI-DIs (wet lab vs. software), and guided Stage I/II audits.
- Why it matters: Ultimately, this demonstrates a repeatable pathway to IVDR certification for large NGS services and software, something that hadno clear precedent.
Read the announcements: For details, read the Fulgent press release and Citeline case study.
0+
Genes Certified
Class C
IVDR Classification
0 mo
To CE Mark
0+
Post-Cert Scale
The challenge: certifying a service-based, large-scale NGS system under IVDR
To begin with, FulgentExome is a service-based NGS solution that integrates wet-lab workflows with the Fulgent PLM bioinformatics pipeline. As a result, pursuing IVDR certification meant converting a mature CLIA/CAP testing service into a CE-marked IVD system with robust evidence across scientific validity, analytical performance, and clinical performance, for thousands of genes. In particular, key hurdles included: defining intended use at scale; validating third-party components; proving scientific validity across 4,600+ genes; above all fully validating the bioinformatics pipeline under medical device software standards.
MDx approach: a playbook for complex NGS + software
1) Build the right QMS, fast
First, we performed an IVDR GAP assessment. Next, we designed and implemented an ISO 13485-compliant QMS with risk management, supplier control, document control, internal audits, and management review—migrating from a CLIA/CAP model to IVDR-ready operations.
2) Engineer a defensible intended use
Meanwhile, the intended-use statement evolved iteratively, from an initial ~300-gene scope to whole-exome, finally landing on 4,600+ genes aligned to available clinical and analytical evidence. In the end, the final language was future-proofed to support rapid updates as evidence expands.
3) Split wet lab and software into two regulated products
Afterward, following round 1 review feedback, we separated the documentation into two Basic UDI-DIs, FulgentExome (wet lab) and Fulgent PLM (software) to align with IVDR expectations for traceability and lifecycle control. Consequently, this restructuring sharpened conformity assessment and accelerated subsequent approvals.
4) Validate the informatics stack like a medical device
In parallel, we validated PLM under IEC 62304/82304, including architecture, version control, cybersecurity, verification/validation, and integration with external databases. Therefore, the result was a fully evidence-backed bioinformatics pipeline capable of reproducible, high-confidence variant calling and reporting.
5) Make “evidence at scale” practical
- First, Scientific validity: Three-tier strategy combining validation of exome sequencing as an approach, reliance on curated public databases, and deep exemplars for a large subset of genes.
- Second, Clinical performance: Leveraged routine testing experience (thousands of positives) to focus clinical evidence on high-prevalence genes, and instituted a robust PMPF strategy to continuously strengthen low-prevalence areas.
6) Orchestrate TÜV SÜD audits to success
- Initially, Stage I confirmed documentation readiness, scope, Basic UDI-DIs and integration of IVDR processes into daily practice.
- Subsequently, Stage II verified on-the-floor implementation of SOPs, training, competence, CAPA and change control—closing findings on short cycles to hit NB timelines.
Results that move the market
- CE mark granted for FulgentExome & Fulgent PLM, among the first end-to-end Class C germline NGS solutions under IVDR.
- Certified scope covers 4,600+ genes, positioning Fulgent as a reference lab for comprehensive hereditary disease testing serving European patients.
- Post-certification, the platform scaled to >7,000 genes using a new probe set, demonstrating the inherent scalability built into the certified system (process, documentation, and change control).
- Strengthened competitive standing in the EU diagnostics market; public communications highlight the magnitude of this achievement for large NGS panels.
Read more in the Fulgent press release and Citeline’s in-depth article.
What this means for labs and IVD developers planning large NGS submissions
If you operate an LDT today: you’ll need to translate CLIA/15189 practices into an ISO 13485 framework, document design controls, and produce a full PER (PEP/PER), APR, SVR, PMS/PMPF, SSP, and labeling/IFU aligned to GSPR. Expect to validate any bioinformatics pipeline as SaMD with IEC 62304/82304 and cybersecurity controls.
If your panel is “large”: you likely won’t have uniform clinical data across every gene. A structured tiered evidence model + PMPF can satisfy Notified Bodies while keeping your roadmap scalable.
If you combine wet lab + software: plan for separate Basic UDI-DIs and documentation sets. Treat the pipeline as a product with its own requirements, verification, and risk controls.
Why MDx
- End-to-end IVDR expertise: From regulatory strategy & classification to Annex II/III technical files, PER/SVR/APR, training, and mock NB reviews. Read more about our NGS regulatory services.
- Clinical performance studies: We design, run, and report ISO 20916 studies (protocols, eTMF, monitoring, biostats, PER alignment), and we can act as delegated sponsor for multi-country submissions—100% submission success rate to date.
- Operational scale: ISO 9001 clinical QMS, EU/US partner network, multilingual CRAs, and a repeatable process honed on 60+ performance study submissions for top IVD and pharma clients.
Project timeline
Our joint project with Fulgent spanned July 2023–July 2025, with overlapping tracks for QMS creation, technical documentation, NB review, and Stage I/II audits, a coordinated plan that allowed rapid closure of findings and post-certification scaling.
Client perspective
The program demanded evening/weekend execution across regulatory, documentation, and project management to meet Notified Body timelines, effort that, in the client’s words, “made all the difference” in achieving what initially “seemed almost impossible.”
Planning IVDR for your NGS panel? Here’s a quick readiness checklist
- Intended use aligned to evidence (and future updates)
- ISO 13485 QMS with software lifecycle integration
- PER (PEP/PER), SVR, APR mapped to gene-level strategy
- PLM/DR pipeline validated per IEC 62304/82304 (+cybersecurity)
- Separate documentation/UDI for wet lab vs. software (if applicable)
- PMS/PMPF plan to mature low-prevalence evidence post-market
- Mock NB review + Stage I/II audit readiness
(Our team can lead or co-author each artifact above.)
Talk to us
Whether you’re certifying a focused oncology panel or pushing the limits with exome-scale content, MDx brings the cross-functional regulatory, clinical, quality, and software depth to make it possible—on a timeline that keeps your business competitive.
For a large, complex NGS panel (thousands of genes, wet lab + bioinformatics software), expect 18 to 24 months from project kickoff to CE mark, assuming you need to build a QMS from scratch. If you already have an ISO 13485-certified QMS and partial technical documentation, the timeline can shorten to 12 to 16 months. The main variables are: the scope of the panel (more genes = more validation work), whether the bioinformatics pipeline needs IEC 62304 validation from zero, Notified Body capacity and review cycles, and the maturity of your clinical evidence. In the Fulgent case, the full project spanned 24 months (July 2023 to July 2025), including QMS creation, full Annex II/III technical documentation, and TÜV SÜD Stage I and Stage II audits.
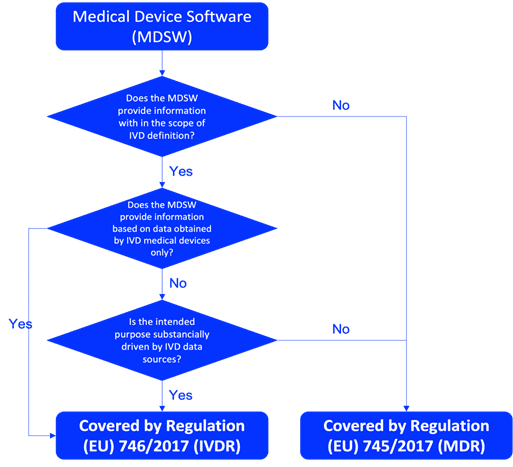
Most NGS-based IVDs classify as IVDR Class C under Annex VIII classification rules, because they typically provide information used to determine patient predisposition or individual risk for serious conditions (e.g., hereditary cancer panels, germline disease testing). NGS panels intended for infectious disease detection with high public health risk (e.g., HIV, hepatitis) may classify as Class D. Companion diagnostic NGS panels co-developed with a therapeutic product also typically fall under Class C. Classification depends on the specific intended use and clinical claims, not the technology itself. All Class C and D IVDs require Notified Body conformity assessment.
Yes, when the bioinformatics pipeline qualifies as standalone software (SaMD) or is a distinct regulated component, IVDR requires a separate Basic UDI-DI. In the Fulgent case, MDx split the documentation into two Basic UDI-DIs: one for FulgentExome (the wet-lab component) and one for Fulgent PLM (the bioinformatics pipeline). This separation aligns with IVDR expectations for traceability, lifecycle control, and independent conformity assessment. Each Basic UDI-DI has its own technical documentation, risk management file, and performance evaluation. This approach also makes post-market updates easier, a software update does not trigger re-review of the entire wet-lab documentation.
No, CLIA/CAP accreditation and ISO 15189 certification are not equivalent to ISO 13485, which is the QMS standard required for IVDR CE marking. While CLIA/CAP provides a strong operational foundation (proficiency testing, personnel qualifications, quality control), it does not cover medical device design controls, supplier management, CAPA, post-market surveillance, or the device lifecycle documentation that IVDR demands. Laboratories transitioning from CLIA/CAP to IVDR must implement an ISO 13485-compliant QMS and document design inputs, outputs, verification, validation, and change control for each IVD product.
For panels targeting thousands of genes, it is typically not feasible to generate individual clinical evidence for every gene-disease association. A tiered approach addresses this: Tier 1 validates the underlying sequencing technology (e.g., exome sequencing as a methodology) with evidence from published literature and peer-reviewed validation studies. Tier 2 relies on curated public databases such as ClinVar, OMIM, and HGMD to establish gene-disease associations at scale. Tier 3 provides deep exemplar evidence (including analytical and clinical performance data) for a representative subset of high-prevalence genes. Genes with limited data are supported through a Post-Market Performance Follow-up (PMPF) plan that progressively strengthens evidence after CE marking. This strategy was accepted by TÜV SÜD in the Fulgent certification.